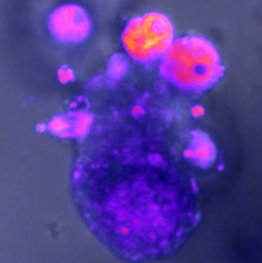
The development of an organism from a single cell and its long-term maintenance require tight coordination of cellular processes at the tissue- and organism-wide level. Signalling pathways have long been studied as key mediators of this control, but only in recent years has it become possible to experimentally investigate the role of signalling dynamics — the changes in signalling activity over time.
Work in single cells has shown that biological information can be encoded in signal dynamics. With new technological advances, we can now study how this dynamic encoding operates at the multicellular level (reviewed in Sonnen and Aulehla 2014).
In our lab, we apply these approaches to investigate dynamic signal encoding during early embryonic development using model systems such as gastruloids, blastoids, and blastocysts. We also study signalling dynamics in tissue homeostasis and regeneration, with a particular focus on the intestine, and explore how disruptions in these processes can drive cancer.
Our approach to study of signalling dynamics
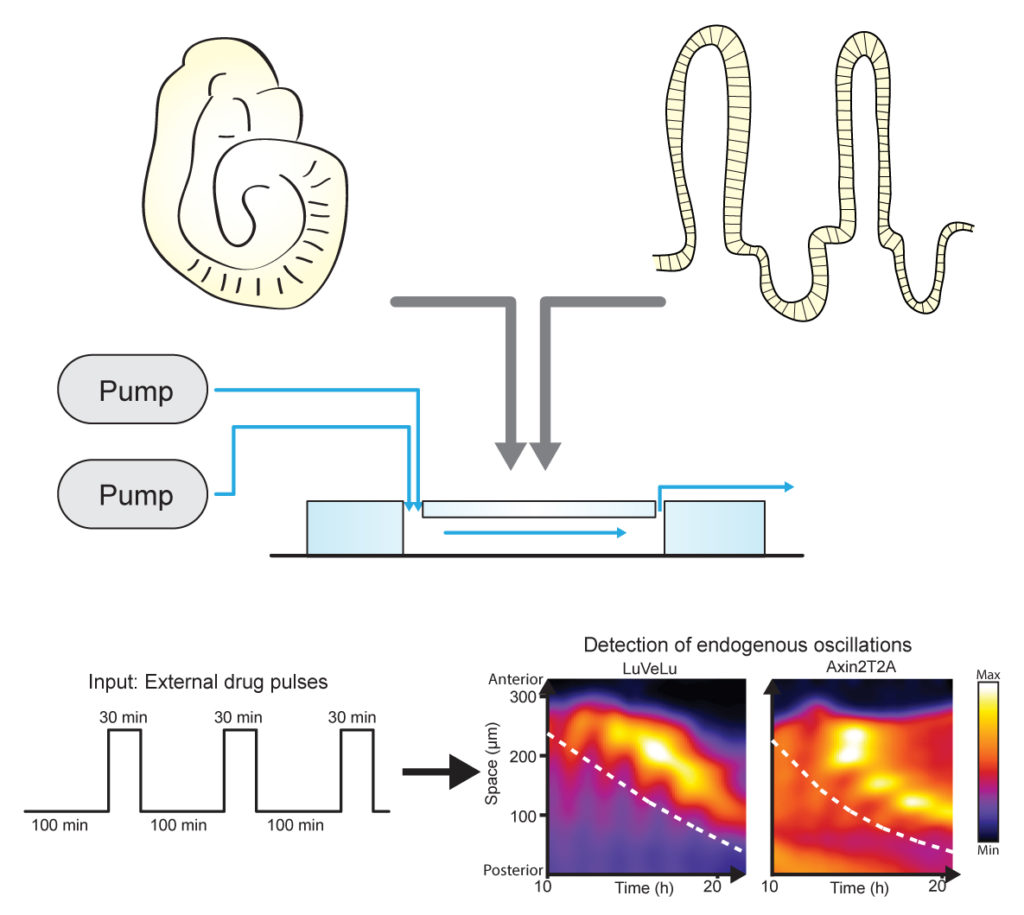
We take a multidisciplinary approach that combines developmental biology, biochemistry, and cell biology with quantitative tools to investigate the function and mechanism of dynamic signal encoding.
To quantify signalling dynamics at cellular and tissue-wide levels, we perform real-time fluorescence imaging of dynamic signalling reporters in model systems such as gastruloids, blastoids, and organoids. These approaches allow us to study signalling during early embryonic development, tissue regeneration, and cancer progression. In addition, we integrate time-resolved proteomics (preprint) and single-cell tracking (preprint) to achieve high-resolution, dynamic measurements of signalling networks.
For functional investigations, we have established a microfluidic system (Sonnen et al. 2018, Sonnen and Merten 2019) that enables precise control of signalling dynamics by applying external pathway modulators with high spatiotemporal resolution. Using this technology, we can manipulate the period of signalling oscillations or control the phase relationship between multiple oscillatory pathways. Beyond microfluidics, we also employ complementary tools such as optogenetics to perturb signalling with equally high spatiotemporal precision.
(figure reprinted and modified from Cell according to CC BY-NC-ND 4.0, Sonnen et al. 2018)
Development and optimization of in vitro model systems
To enable precise experimental manipulation and to reduce reliance on animal models, we focus on developing and optimizing in vitro model systems. These systems provide powerful platforms to study both embryonic development and adult tissue biology under controlled conditions. Over the last years, the field has established diverse models, such as the in vitro differentiation of embryonic stem cells, gastruloids, and organoids.
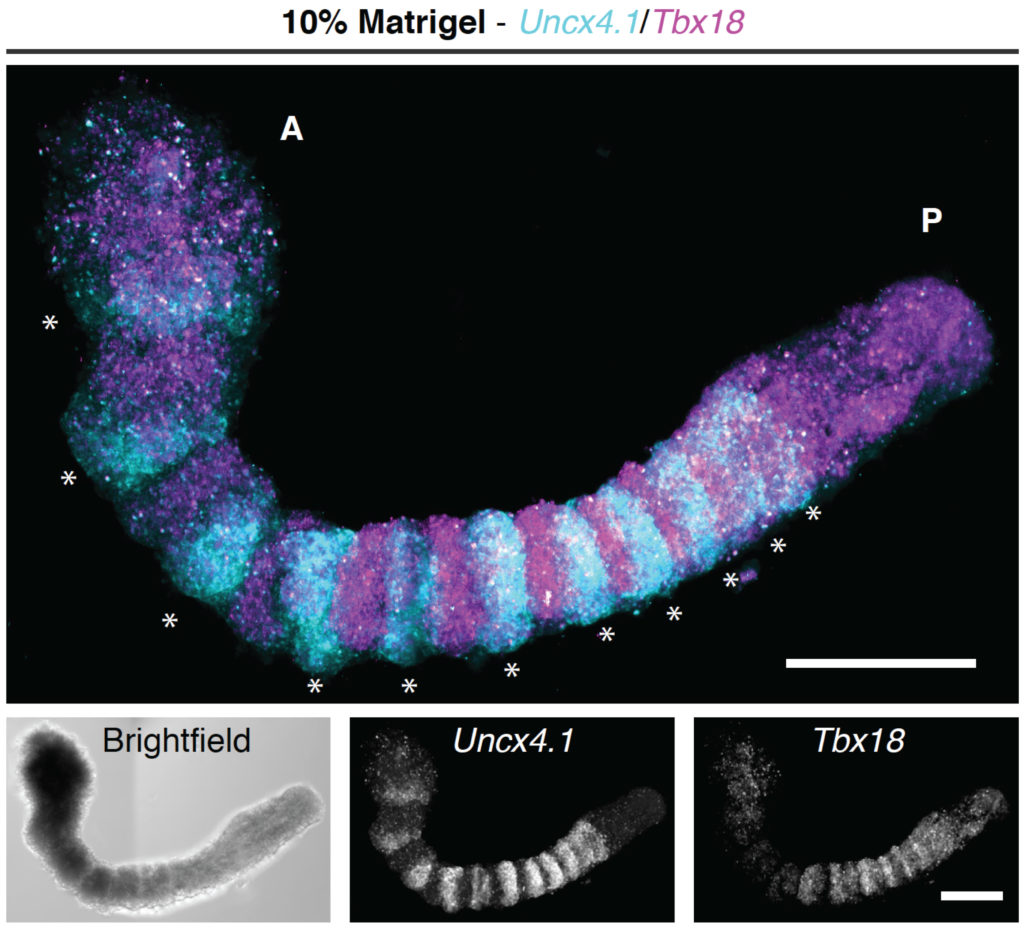
In our lab, we have recently worked on establishing gastruloids as a model for somitogenesis — the periodic segmentation process of the vertebrate embryo — in collaboration with the groups of Alfonso Martinez Arias and Alexander van Oudenaarden. Using these in vitro structures, we demonstrated the presence of signalling oscillations and optimized culture conditions to allow for proper segment formation (van den Brink et al. 2020, Batenburg et al. 2020).
(figure reprinted and modified from Nature, van den Brink et al. 2020)
How do signalling dynamics control embryonic development?
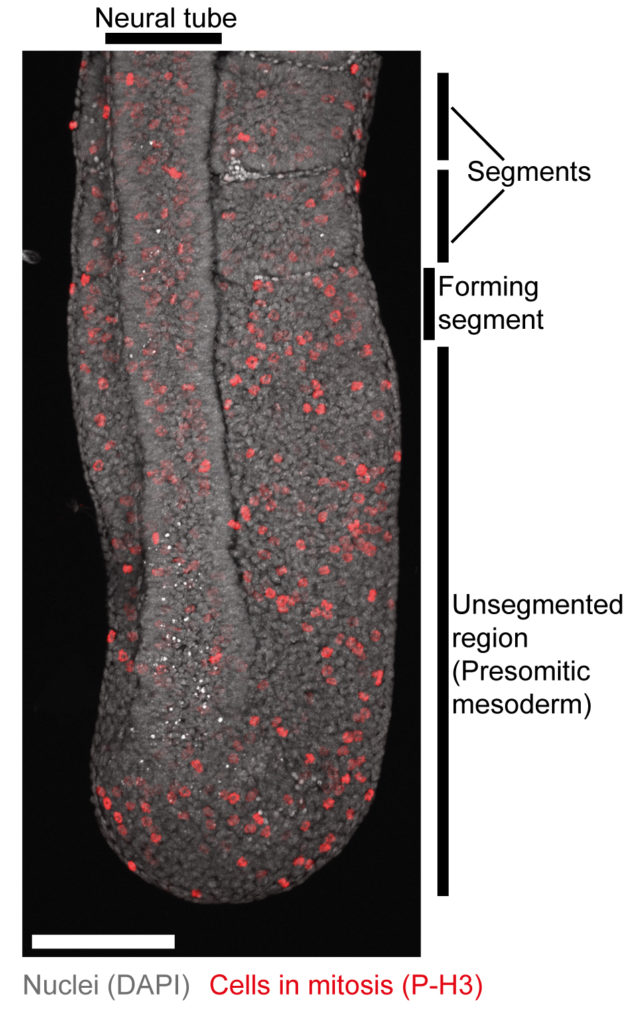
Somitogenesis is the periodic formation of somites — the building blocks of vertebrae and axial muscles — during early embryonic development. This sequential segmentation of the presomitic mesoderm (PSM) is regulated by a combination of signalling gradients and oscillations. Oscillatory activity in the Notch, Wnt, and FGF pathways forms the so-called segmentation clock, which determines the timing of somite formation.
Our work has shown that critical information for periodic segmentation is encoded not simply in the oscillations themselves, but in the relative timing between Wnt and Notch signalling oscillations (Sonnen et al. 2018).
To investigate these mechanisms, we use both in vivo models and advanced in vitro systems. For early development, we study blastocysts and blastoids to explore how signalling dynamics guide the earliest steps of embryogenesis, while gastruloids allow us to model complex processes such as somitogenesis.
Together, these approaches help us uncover how dynamic signalling controls tissue patterning and timing during embryonic development, and provide insights into how disruptions in these processes may contribute to disease.

How do signalling dynamics control adult tissue homeostasis?

In adult tissues, signalling pathways regulate cell turnover and differentiation to maintain homeostasis. A well-studied example is the small intestine, where key pathways controlling stem cell renewal and differentiation have been identified. Yet, the role of signalling dynamics in these processes is still largely unknown.
The development of organoid cultures now allows us to grow adult tissues ex vivo, providing a controlled system to study dynamic regulation. By combining organoids with dynamic signalling reporters, real-time imaging, and microfluidic-based perturbations, we can dissect how signalling dynamics govern tissue homeostasis and regeneration.
Our recent preprint demonstrates that these tools allow precise monitoring and manipulation of signalling pathways in organoid cultures, revealing how the timing and coordination of signals control tissue maintenance and how disruptions can contribute to cancer.

How do signalling dynamics impact on disease development and treatment?
Signalling pathways regulate a wide range of processes in the body, from embryonic development to tissue regeneration and homeostasis. Mutations or misregulation of these pathways can lead to developmental disorders or diseases such as cancer.
In our lab, we study how changes in signalling dynamics contribute to cancer development and explore whether these dynamic properties can be targeted for therapy. With support from the KWF Dutch Cancer Society, we are investigating how alterations in signalling timing and coordination drive tumor formation and progression. By combining advanced imaging, organoid models, and precise pathway perturbations, we aim to uncover new strategies to intervene in cancer at the level of temporal and spatial signal control.